 ODAY I AM releasing the final code from my current project. The following is the accompanying
ODAY I AM releasing the final code from my current project. The following is the accompanying README file:
Homepage: http://schestowitz.com/Research/Cardiac/Cardiomat
Index of related documents: http://schestowitz.com/Research/Progress/2010-2011.htm
INTRODUCTION
Cardiomat is the name given to an Octave/MATLAB program whose intended use is heart tracking, with various different bits that help probe cardiac structure and perform basic measurements.
FILES
Contained in the package are the following M-files and sub-directories:
./Experiments
./Experiments/fiesta_simulation-old-experiments.m
./create_synthetic_set.m
./GUI
./GUI/cardiomat.png
./GUI/splash.m
./GUI/splash.fig
./GUI/get_parameters_from_user.m
./GUI/getoptionsgui.m
./GUI/cardiomat-title.jpg
./GUI/cardiomat.m
./GUI/cardiomat.jpg
./getdicomimages.m
./loadtaggedsequence.m
./README (this file)
./Imported
./Imported/findit
./Imported/findit/findit.m
./Imported/arrow
./Imported/arrow/arrow.m
./Imported/gradient
./Imported/gradient/gaussgradient.zip
./Imported/gradient/gaussgradient
./Imported/gradient/gaussgradient/gaussgradient.m
./Imported/gradient/gaussgradient/README.txt.txt
./Imported/gradient/gaussgradient/testgaussgradient.m
./Hybrid
./Hybrid/cardsim_frame_shortaxis.m
./Hybrid/shuffle_transform.m
./fiesta_simulation.m
In addition, the program depends on at least one function from biosig4octmat-2.50, whose homepage is located at http://biosig.sourceforge.net. That function helps find the centre and the radius of a best fit circle/sphere for a group of points.
The directory named “Hybrid” contains functions that were written jointly and “Imported” contains functions which were brought from the outside. The core functions in the package are GPLv3-licensed and the rest are most likely to be BSD-licensed (check the sources to verify). Some of the imported functions may work only in MATLAB and not in GNU Octave, which is a free/libre substitute for MATLAB that lacks some graphical features for the most part.
HOW TO RUN THE PROGRAM
The program was written to be compatible with both Octave and MATLAB. There are some features in it that may require a UNIX/Linux system (e.g. animated videos) and some that may require MATLAB. Additionally, in order to reduce dependency on MATLAB’s GUI utilities, the program can be run in GUI and CLI modes separately, with only the use of figures being the exception to this (figures are output rather than means of interaction).
Key functions reside in the main directory of the program (one function, getdicomimages.m, is deprecated) and a directory named “GUI” contains some of the newer parts that add a GUI.
An important interface function is called fiesta_simulation.m. The first line of code in this function is a boolean flag that says whether the program should be run in GUI mode or not. If use_gui is 0, then the program will simply run the list of tasks specified below. It is possible to give several jobs for the program to complete this way, e.g. in order to run many experiments overnight. If use_gui is non-zero (probably 1 but not necessarily), then the function is assumed to have been called from the file GUI/cardiomat.m, which is a wrapper that invokes a splash screen and some dialogues that serve as a graphical interface. In turn, parameters fed to these dialogues will be passed to fiesta_simulation.m. It might be necessary to modify some paths inside the files in case the functions and the files are located in different places than the ones specified in parameters. Such paths are named in fiesta_simulation.m, loadtaggedsequence.m (specifies output directories onto which to save figures), and splash.m (looks for program images). For a new user, the only things require modifications are likely to be directory paths and maybe the use_gui boolean, depending on this user’s preference.
To run the program in GUI mode, MATLAB is needed. There is also a parameter to specify in fiesta_simulation.m in order to inform loadtaggedsequence.m whether MATLAB or Octave gets used. It is the boolean which is also the last argument in the function call, being 0 for MATLAB and 1 for Octave. The GUI front end will always pass 0 under the assumption that it is run under MATLAB.
DATASETS
The specified directories include a path from which to open images, optionally with a wildcard to filter bits contained within a given directory, e.g. selecting a subset of images that are a range of numbers. Due to privacy/data anonymity constraints, no raw data accompanies this program, even though it can definitely be obtained from many places (see for example https://schestowitz.com/Weblog/archives/2010/11/14/segmented-and-tagged-datasets/).
COMPATIBILITY
There are separate levels of compatibility dilemmas. One is the program level and another is the operating system level. Although the program has been tested in both MATLAB and Octave, on both GNU/Linux and Windows, there may be trivial code changes to make in case errors show up. Most will be path related, so it is abundantly clear that emphasis should be put on making all functions reachable from within paths definition (e.g. “addpath(genpath(YOUR_PROGRAM_ROOT_DIR_HERE))”) and booleans should be set to do the right thing (GUI/CLI and MATLAB/Octave). The goal is to ensure the program can be run under as many platforms as possible, with either free/libre or proprietary software.
IN CASE OF DIFFICULTIES
If issues running the program are experienced, please contact the author (address above). Bugs are expected to exist (it is a research purposes program, so polish is not a priority) and input is not validated or sanitised to provide virtual ‘rails’ on which the user operates the program (sane values need to be given). Sample function calls are given in fiesta_simulation.m and /Experiments/fiesta_simulation-old-experiments.m. Some of them may use the old interface and thus lack some input arguments, but most of them show the type of experiments which were run using the program.


 HIS post provides some information of interest to those who may find themselves working with 70 GB of data and some programs [1, 2]. The latter is a FRGC Web site. The package comes with associated applications and scripts written in Java, C++, Perl, etc. The previous post about the dataset (FRGC ver2.0) offers a bit of background that is research-specific (relating to Dr. Ajmal Mian and his Ph.D. student Faisal R. Al-Osaimi), whereas the notes below are a bit more generic. This series of posts is not about statistical models of faces but only about the dataset. This recent message from Face Recognition Research Community contains MATLAB/Octave loader code for a data instance from the dataset, where each 3-D face weighs about 13 MB (compressed):
HIS post provides some information of interest to those who may find themselves working with 70 GB of data and some programs [1, 2]. The latter is a FRGC Web site. The package comes with associated applications and scripts written in Java, C++, Perl, etc. The previous post about the dataset (FRGC ver2.0) offers a bit of background that is research-specific (relating to Dr. Ajmal Mian and his Ph.D. student Faisal R. Al-Osaimi), whereas the notes below are a bit more generic. This series of posts is not about statistical models of faces but only about the dataset. This recent message from Face Recognition Research Community contains MATLAB/Octave loader code for a data instance from the dataset, where each 3-D face weighs about 13 MB (compressed):





 Filed under:
Filed under: 

 ver the past few days I’ve been repackaging one of my programs such that it becomes easier to install and more user-friendly once installed. The program works in both MATLAB (proprietary software) and Octave (Free/libre software) and it now has a GUI and a name, which I arbitrarily and hastily named “Cardiomat”. Directory selection dialogues and some other user interface features are missing from Octave, so the code has become filled with some conditional statements so that it serves each user, no matter the choice of interpreters (proprietary or Free).
ver the past few days I’ve been repackaging one of my programs such that it becomes easier to install and more user-friendly once installed. The program works in both MATLAB (proprietary software) and Octave (Free/libre software) and it now has a GUI and a name, which I arbitrarily and hastily named “Cardiomat”. Directory selection dialogues and some other user interface features are missing from Octave, so the code has become filled with some conditional statements so that it serves each user, no matter the choice of interpreters (proprietary or Free).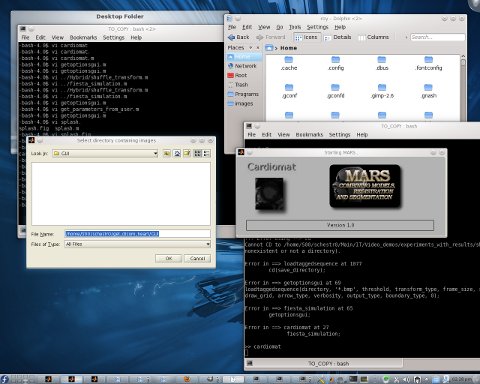


 S a first validation step I am running these basic sanity checks which track anatomy where the correct correspondences are known. These animations show how a collection of points spread around the boundary of a circle (but not put precisely on the circle, on purpose) move along with the image merely by tracking the signal, without any knowledge of what other points are doing or what rigid/affine transformation the image itself is subjected to. Moreover, the interpolation process causes little issue at all and even though points can only move up/down/left/right (and diagonally), they do manage to move consistently.
S a first validation step I am running these basic sanity checks which track anatomy where the correct correspondences are known. These animations show how a collection of points spread around the boundary of a circle (but not put precisely on the circle, on purpose) move along with the image merely by tracking the signal, without any knowledge of what other points are doing or what rigid/affine transformation the image itself is subjected to. Moreover, the interpolation process causes little issue at all and even though points can only move up/down/left/right (and diagonally), they do manage to move consistently.



 ORKING with bare metal for the purpose of maximal hardware utilisation and performance is not the same as working with typical computer programs. Developing software for research is also different from software development for end users, where the software is treated as a product. Different scenarios require different methodologies and different levels of polish, and thus have different specifications and priorities. In research, brute force becomes essential to the success of one group or another.
ORKING with bare metal for the purpose of maximal hardware utilisation and performance is not the same as working with typical computer programs. Developing software for research is also different from software development for end users, where the software is treated as a product. Different scenarios require different methodologies and different levels of polish, and thus have different specifications and priorities. In research, brute force becomes essential to the success of one group or another.