 T HAS been a week or so since I last made a research-related post, so here is a quick update. I am currently coding some bits that can detect and measure rotation in the heart, including tangential/centrifugal components. The goal is to calculate the twist and also the scale changes separately, giving a figure indicative of each. The notions found in flow/vortex mathematics ought to enable this to be calculated from a set (or matrix) of points, each of which being a vector of directions around a central point. To quote Wikipedia on vortex: “A vortex (plural: vortices) is a spinning, often turbulent, flow of fluid. Any spiral motion with closed streamlines is vortex flow. The motion of the fluid swirling rapidly around a center is called a vortex. The speed and rate of rotation of the fluid in a free (irrotational) vortex are greatest at the center, and decrease progressively with distance from the center, whereas the speed of a forced (rotational) vortex is zero at the center and increases proportional to the distance from the center. Both types of vortices exhibit a pressure minimum at the center, though the pressure minimum in a free vortex is much lower.”
T HAS been a week or so since I last made a research-related post, so here is a quick update. I am currently coding some bits that can detect and measure rotation in the heart, including tangential/centrifugal components. The goal is to calculate the twist and also the scale changes separately, giving a figure indicative of each. The notions found in flow/vortex mathematics ought to enable this to be calculated from a set (or matrix) of points, each of which being a vector of directions around a central point. To quote Wikipedia on vortex: “A vortex (plural: vortices) is a spinning, often turbulent, flow of fluid. Any spiral motion with closed streamlines is vortex flow. The motion of the fluid swirling rapidly around a center is called a vortex. The speed and rate of rotation of the fluid in a free (irrotational) vortex are greatest at the center, and decrease progressively with distance from the center, whereas the speed of a forced (rotational) vortex is zero at the center and increases proportional to the distance from the center. Both types of vortices exhibit a pressure minimum at the center, though the pressure minimum in a free vortex is much lower.”
My implementation of this is simplistic as it merely uses an accumulator whose value increases for each point that moves in a given direction (e.g. for each clockwise arrow) and decreases for the opposite.
An issue we face here is that circular motion in the heart walls is far from trivial to see. A careful look at image sequences taken with about 20 frames for each cycle and each slice does not quite expose a ‘squeeze’ of the heart, not unless there is tagging of the image which also enables reconstruction of a higher quality. So, to calculate rotation of the sides based on just intensity is rather hard, unless there is something in the signal that can be detected. Generally, looking at the tracking typically carried out, there is hardly a tendency to pick up any traces of tangential motion and where it does exist it is not consistent. Assuming a direction is known and then estimated with a probabilistic model, there is room for improvement, maybe even a significant change in contrast. Different slices show different directions for the twist and it requires a look inside the heart too, in order to visually compare intensities and assume a rotation of some sort. I will post some images soon to illustrate this.
Additional code was completed to calculate and plot the centre of the points around the heart, based on the fitting of a circle to all the surrounding points (best match). At the end of each tracking sequence a plot if shown to give a statistical analysis of how points move around the walls and whether they move in and out throughout the cardiac cycle. The goal here is to, based on several cycle, find out if and how the algorithm can detect wall motion and quantify it. There is clearly a correlation to be found here.
Today’s addition (it’s a holiday, so lots of spare time for coding and also to read literature) will soon be uploaded along with more material, all of which will be shared not just for the sake of transparency. I’ll post results shortly.
As a side note, I’ve begun looking into the possibility of writing a book about Novell and another possibility I’m looking into is initiating a startup around the code I’ve been writing since 2003 to achieve something nobody achieved before, even with free/libre software (the whole stack). I’m currently doing cardiac analysis but still considering a situation where come back to a more CS-oriented work, at least when the existing contract is over. There are many projects on my mind and choosing just the best one/s is not easy. Then again, it’s good to have possibilities and I’m still in my twenties, so there is no pressure.
 currently put the final touches on my program which detects and analyses the movement of the heart using the fiesta protocol in an MRI-type modality (1.5 Tesla).
currently put the final touches on my program which detects and analyses the movement of the heart using the fiesta protocol in an MRI-type modality (1.5 Tesla).










 Filed under:
Filed under: 

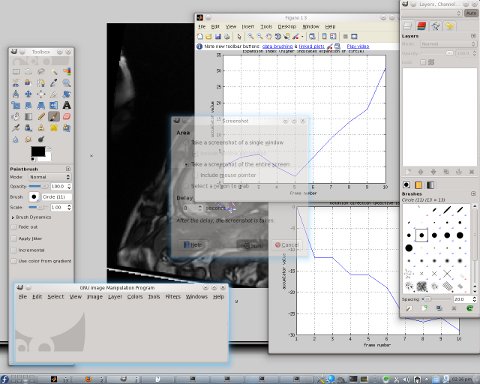
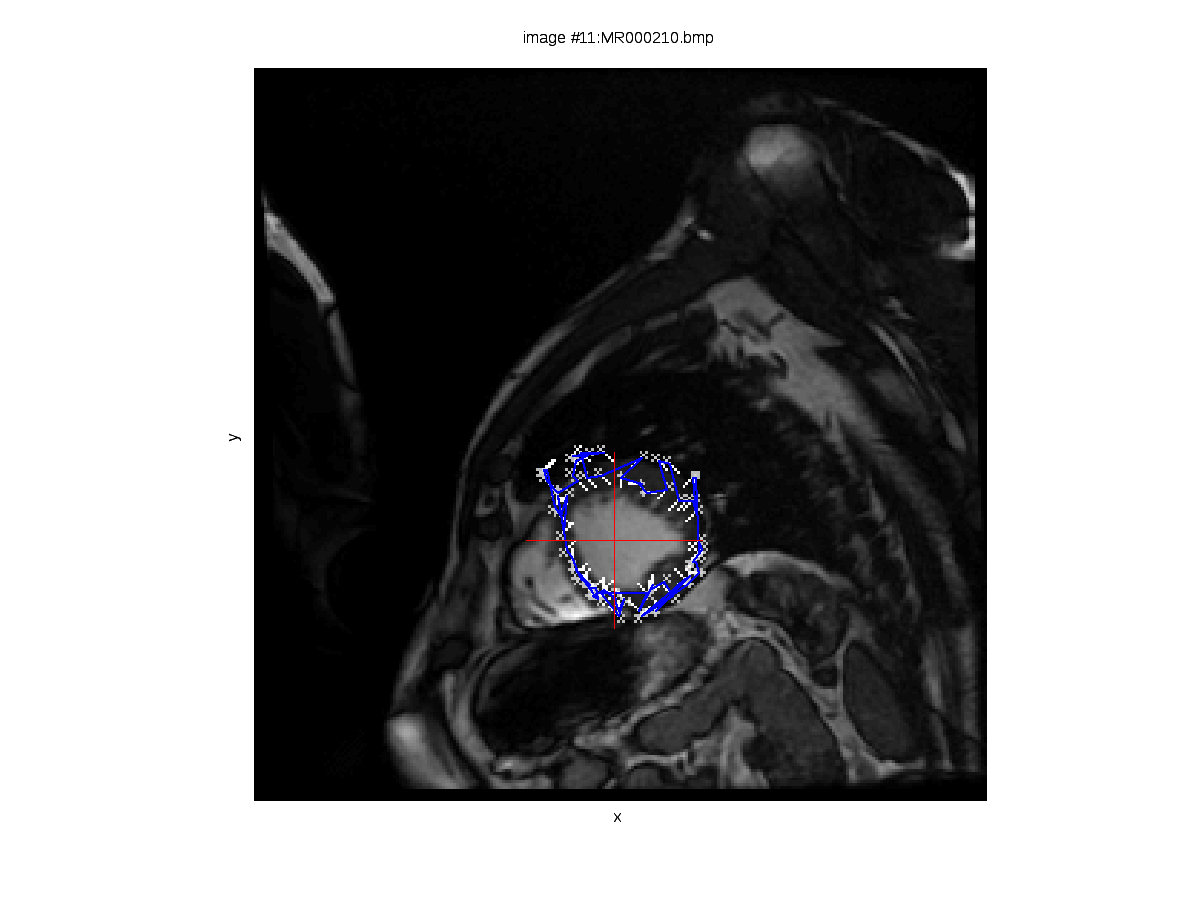
 hown here are two images derived from experiments out of which there are also videos. The set of cardiac images contains 340 images, divided into 17 groups of 20 images belonging to each. A group of 20 consecutive images represents one slice imaged throughout one cardiac cycle. This means that from the 2nd and 4th slides — images 21 and 61 respectively for example — the signal is approximately correspondent and thus can be compared. A series of experiments was performed in turn, dealing with each slice in isolation and learning how our algorithm copes with the complicated task of tracking the heart’s walls without a priori knowledge such as a model of the heart, for example (a model whose parameter values can be optimised over, for a good fit to be eventually found). We are interested in the rotation of the heart at the different vertical levels, particularly because we expect to see clockwise and counterclockwise movements throughout the cycle, depending on the slice (the heart squeezes blood by moving in different — almost opposite — directions).
hown here are two images derived from experiments out of which there are also videos. The set of cardiac images contains 340 images, divided into 17 groups of 20 images belonging to each. A group of 20 consecutive images represents one slice imaged throughout one cardiac cycle. This means that from the 2nd and 4th slides — images 21 and 61 respectively for example — the signal is approximately correspondent and thus can be compared. A series of experiments was performed in turn, dealing with each slice in isolation and learning how our algorithm copes with the complicated task of tracking the heart’s walls without a priori knowledge such as a model of the heart, for example (a model whose parameter values can be optimised over, for a good fit to be eventually found). We are interested in the rotation of the heart at the different vertical levels, particularly because we expect to see clockwise and counterclockwise movements throughout the cycle, depending on the slice (the heart squeezes blood by moving in different — almost opposite — directions).

 or the sake of cross-application/framework compatibility, one occasionally needs to alter code until it works everywhere, without the need to keep two (or more) separate codebases. This situation is far from ideal, but then again, not everything works like Java. When colleagues use a different platform and occasionally prefer proprietary software it is only fair to do some extra work catering for it.
or the sake of cross-application/framework compatibility, one occasionally needs to alter code until it works everywhere, without the need to keep two (or more) separate codebases. This situation is far from ideal, but then again, not everything works like Java. When colleagues use a different platform and occasionally prefer proprietary software it is only fair to do some extra work catering for it. uring the holidays I decided to see what else is out there which is already free/libre software like my own work, which thus can be merged for comparative purposes. It would be valuable to have applied to my work some comparison to existing tracking algorithms which deal with cardiac images. A
uring the holidays I decided to see what else is out there which is already free/libre software like my own work, which thus can be merged for comparative purposes. It would be valuable to have applied to my work some comparison to existing tracking algorithms which deal with cardiac images. A 
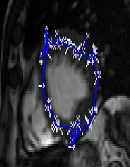
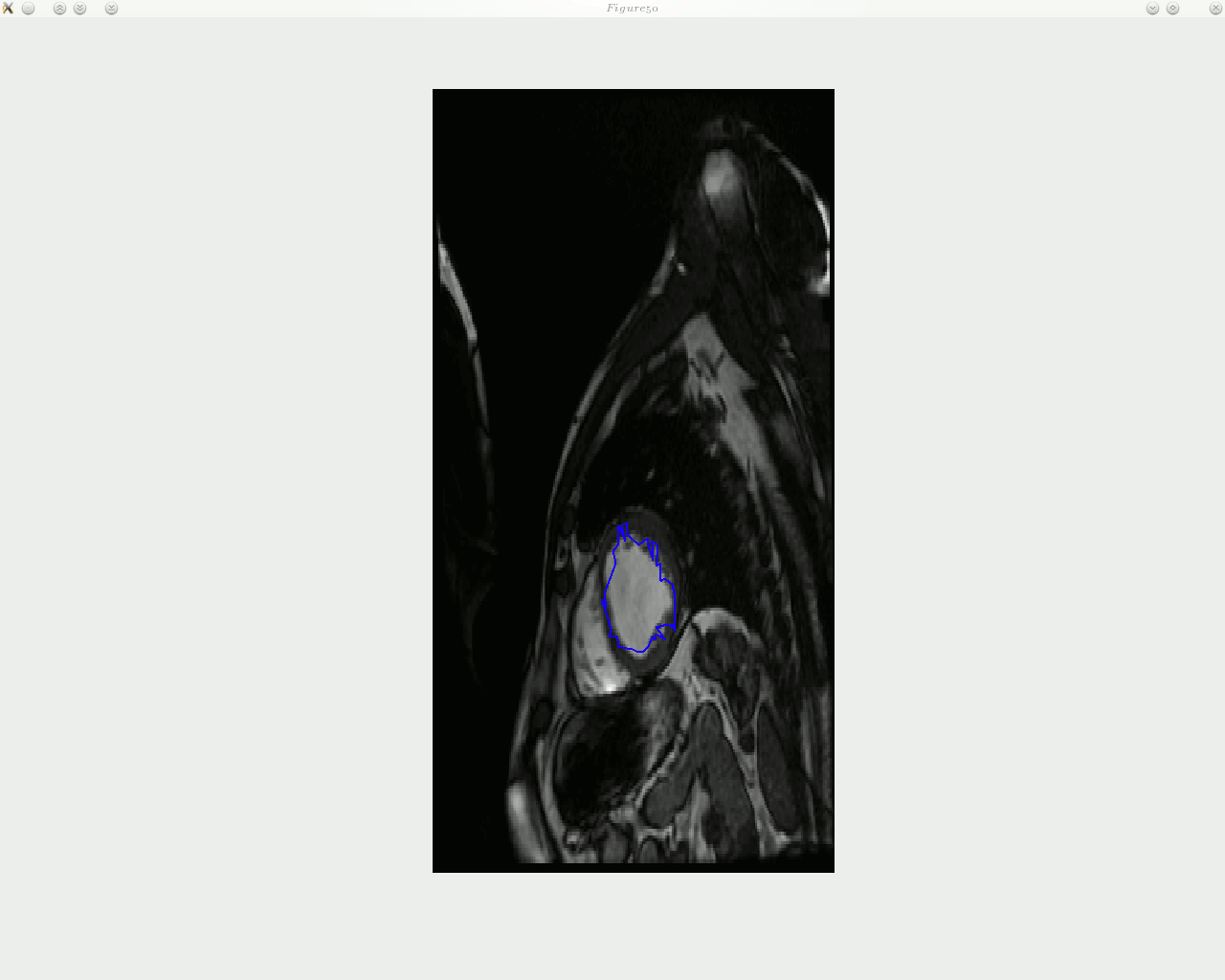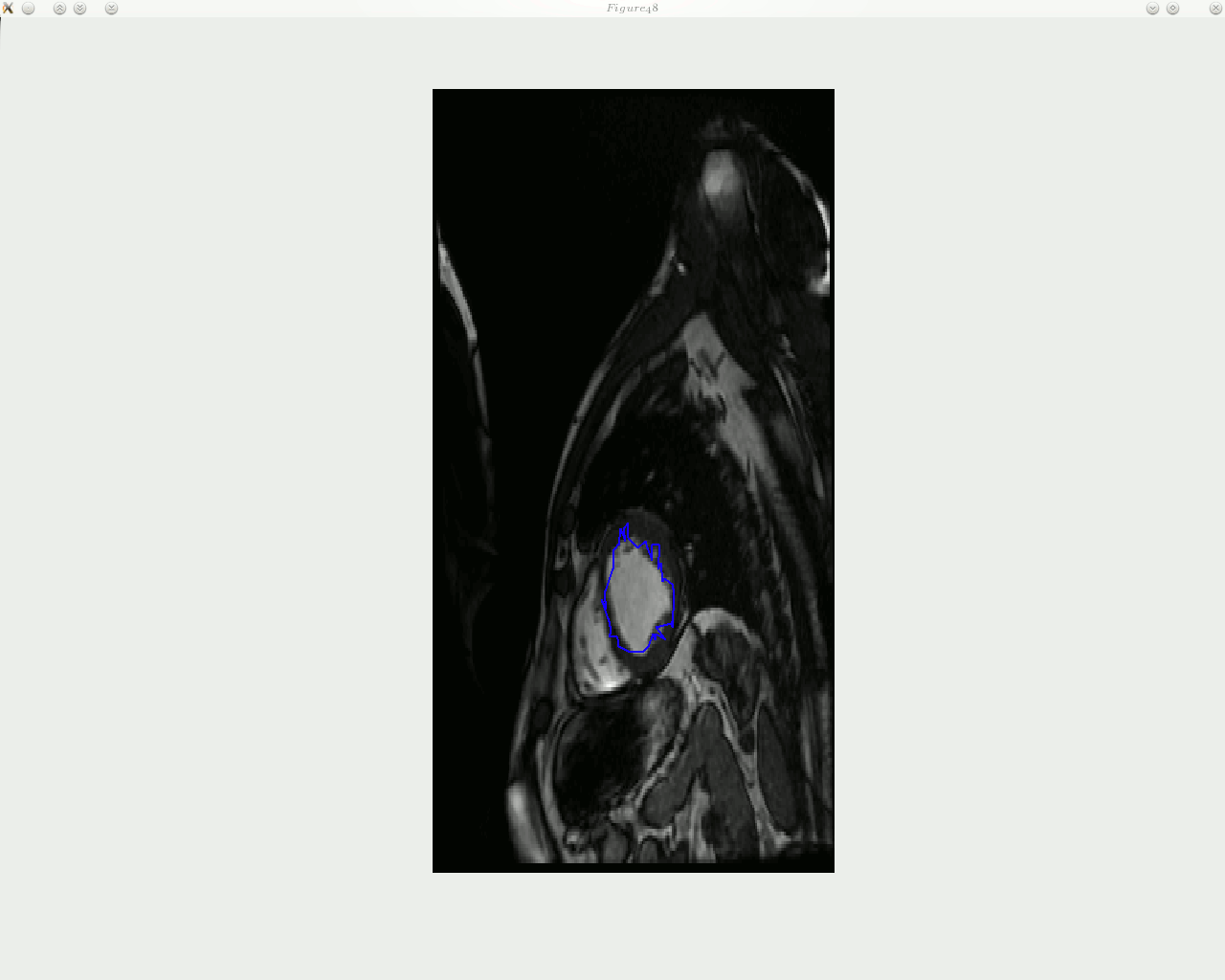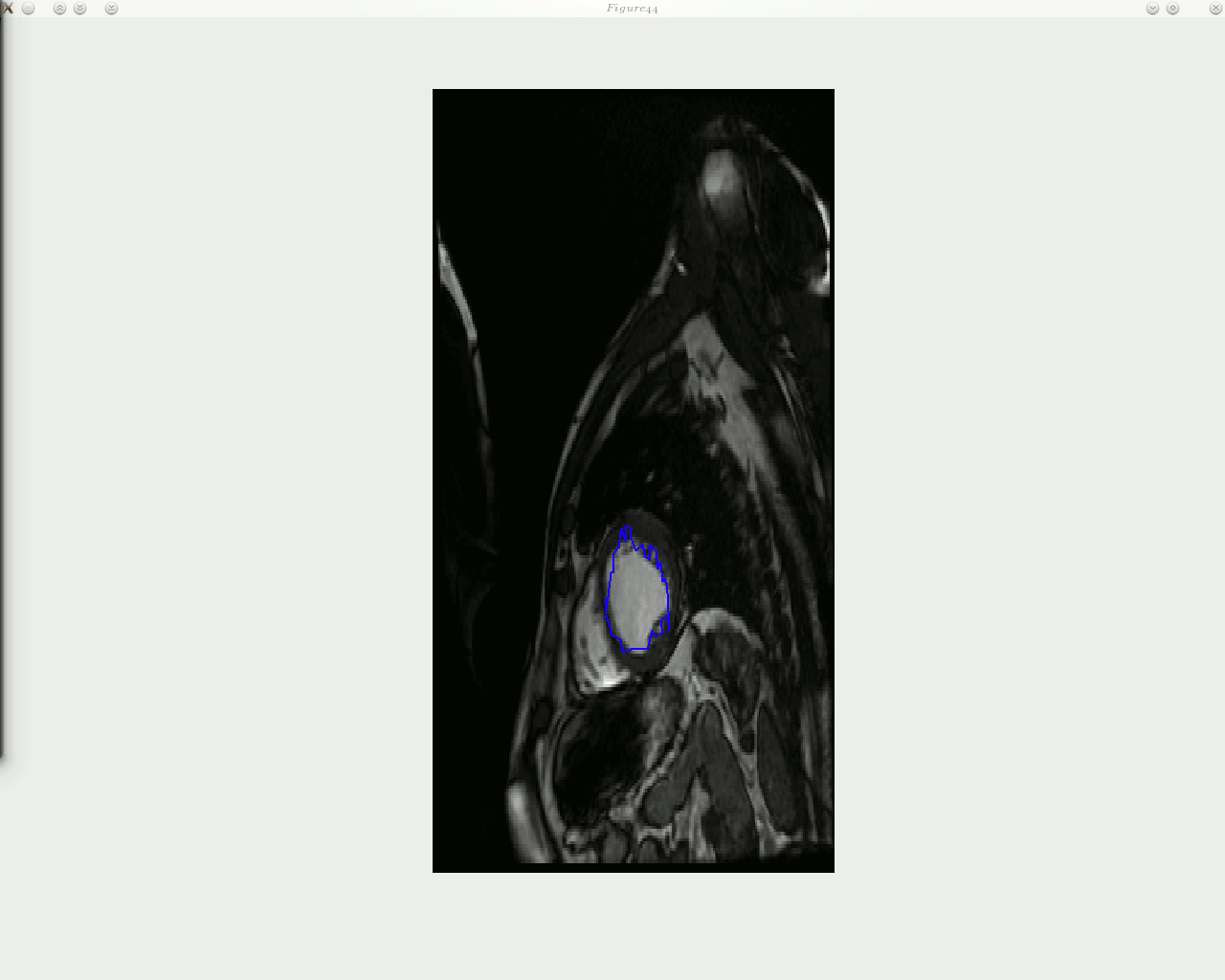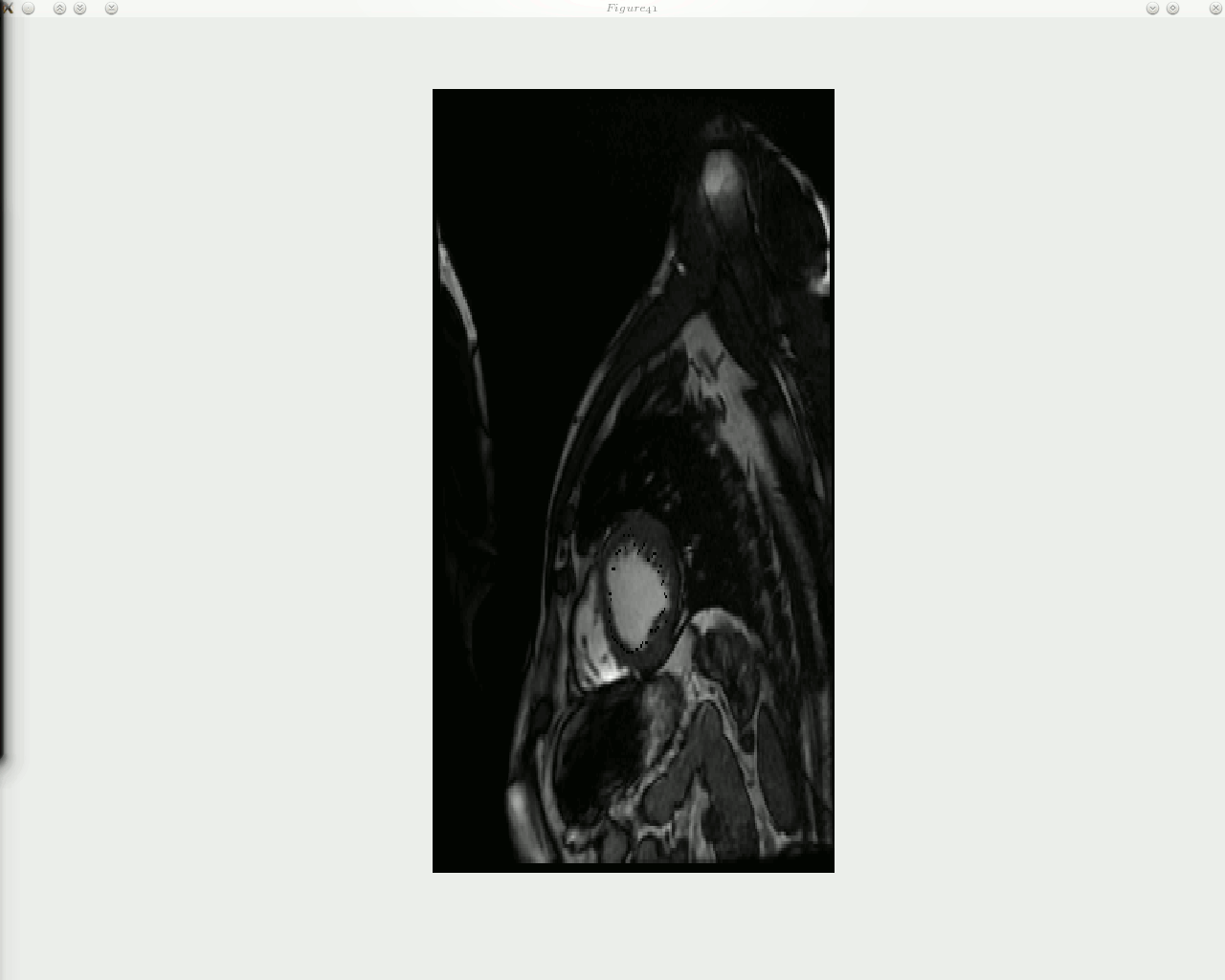
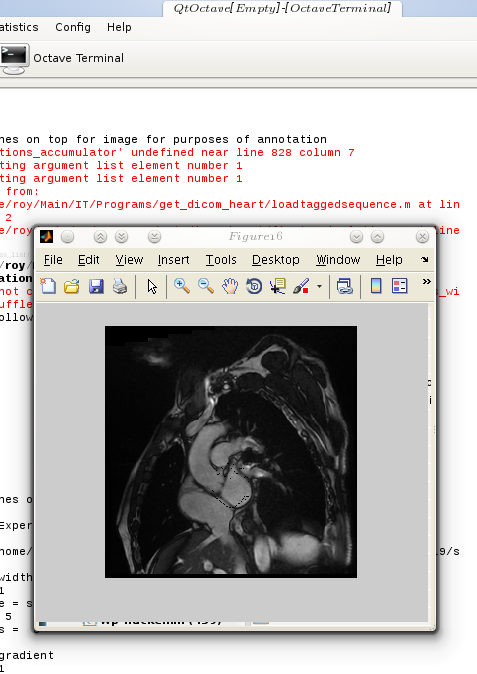
 he animated figure below shows how contouring is done with a new and improved algorithm in place. The sequence is show in reverse, as the window titles help indicate. Arrows can be put on top of the contour to signal the direction of movement of each landmark point, but arrow can in general add clutter and it may be worth taking a group of several points and study based on these the direction of movement of entire region, then display fewer large arrows.
he animated figure below shows how contouring is done with a new and improved algorithm in place. The sequence is show in reverse, as the window titles help indicate. Arrows can be put on top of the contour to signal the direction of movement of each landmark point, but arrow can in general add clutter and it may be worth taking a group of several points and study based on these the direction of movement of entire region, then display fewer large arrows.